3.1.1
Prepare suspensions of ampicillin-sensitive
and -resistant E. coli with densities of 2 X 10^7
cfu/mL in 5% TSB/saline (cfu = colony forming units).
3.1.2
Prepare 2X SYTOX Green staining solution by diluting
the 5 mM SYTOX Green/DMSO stock solution by a factor
of 1:500 in 5% TSB/saline, yielding a final dye
concentration of 10 µM. To prepare sufficient 2X
staining solution for analysis of 10 samples, add
30 µL of 5 mM SYTOX Green/DMSO to 15 mL of 5% TSB/saline.
3.1.3
Prepare a 200 µg/mL stock solution of ampicillin
in 2X SYTOX Green staining solution (step 3.1.2).
5 mL of this stock solution is sufficient for the
measurements specified in this protocol.
3.1.4
Prepare dilutions of the ampicillin stock solution
to concentrations of 2 µg/mL, 20 µg/mL and 60 µg/mL
by addition of further 2X SYTOX Green staining solution
(step 3.1.2).
3.1.5
Label a set of 5 glass test tubes "Resistant"
and a second set of 5 tubes "Sensitive". Label one
member of each set of tubes with the following designations:
Reagent blank, 1, 10, 30 and 100 µg/mL ampicillin.
3.1.6
Transfer 1.5 mL of TSB/saline/2X SYTOX Green (step
3.1.2) to each reagent blank test tube and make
up to 3 mL total volume by adding a further 1.5
mL TSB/saline.
3.1.7
Transfer 1.5 mL of 2 µg/mL, 20 µg/mL, 60 µg/mL and
200 µg/mL ampicillin in 2X SYTOX Green staining
solution (steps 3.1.3 and 3.1.4) to the tubes labeled
1, 10, 30 and 100 µg/mL ampicillin, respectively.
3.1.8
Transfer 1.5 mL of the resistant E. coli suspension
(step 3.1.1) to each of the 4 ampicillin-containing
test tubes labeled "Resistant".
3.1.9
Transfer 1.5 mL of the sensitive E. coli suspension
(step 3.1.1) to each of the 4 ampicillin-containing
test tubes labeled "Sensitive".
3.1.10
Incubate all samples at 37° C for 1 hour prior to
carrying out fluorescence measurements.
3.2.1
Set-up TD-700 with a quartz halogen lamp, Omega
Optical 485DF22 excitation filter, and Omega Optical
530DF30 emission filter.
3.2.2
Calibrate in the Simple Mode according to section
VII of the TD-700 Manual using the sample from the
antibiotic-sensitive set that contains the highest
concentration of ampicillin (100 µg/mL) as the calibration
standard. This sample should be the most fluorescent
among those to be measured.
3.2.3
Read the "Sensitive" and "Resistant" samples in
the order of reagent blank followed by the bacterial
samples in order of increasing ampicillin concentration.
3.2.4
For each sample, record the fluorescence intensity
value shown on the fluorometer display screen. To
equalize any photobleaching effects, insert samples
into the fluorometer for approximately equal time
periods. The data may be corrected for the dye background
by subtracting the reagent blank fluorescence value
from bacterial sample fluorescence readings.
3.2.5
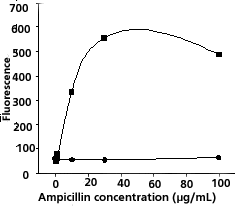
Plotted data should show increasing fluorescence
as a function of ampicillin concentration for antibiotic-sensitive
bacteria in contrast to relatively constant low
fluorescence for antibiotic-resistant bacteria (Figure
1).